Tutorial: Distributed training across multiple nodes.#
Note The provided commands to execute in this tutorial assume include a configuration script to
clone the full library. Setting up headless THOR might
require superuser privileges. We also assume NCCL is available for communication
across computation nodes and all nodes have a running ssh server.
The below introduced experimental tools and commands for distributed training assume a Linux OS (tested on Ubuntu 18.04).
In this tutorial, we:
- Introduce the available API for training across multiple nodes, as well as experimental scripts for distributed configuration, training start and termination, and remote command execution.
- Introduce the headless mode for AI2-THOR in
AllenAct. Note that, in contrast with previous tutorials using AI2-THOR, this time we don't require an xserver (in Linux) to be active. - Show a training example for RoboTHOR ObjectNav on a cluster, with each node having sufficient GPUs and GPU memory to host 60 experience samplers collecting rollout data.
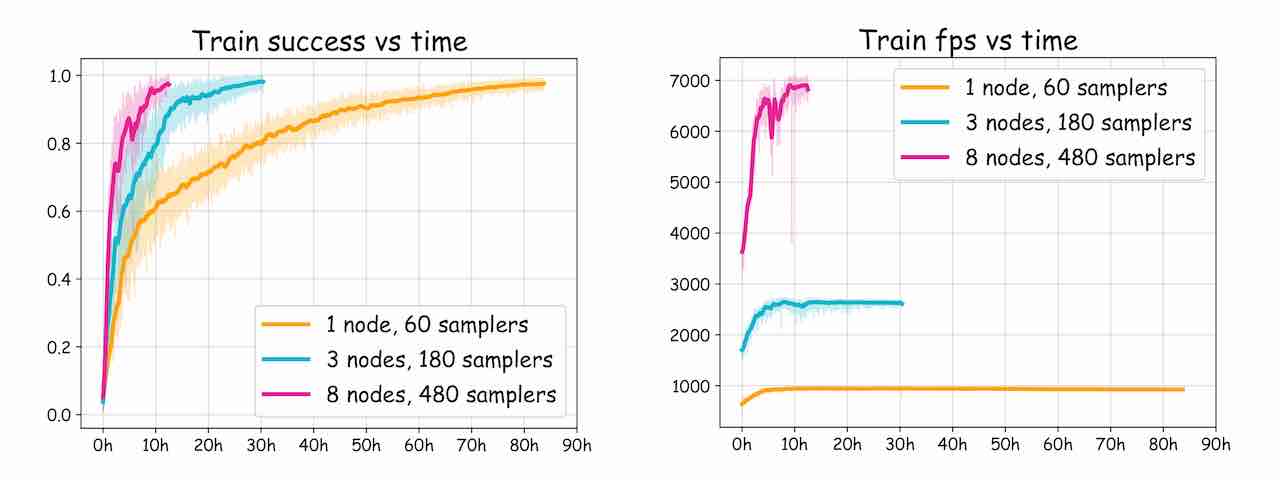
Thanks to the massive parallelization of experience collection and model training enabled by DD-PPO, we can greatly speed up training by scaling across multiple nodes:
The task: ObjectNav#
In ObjectNav, the goal for the agent is to navigate to an object (possibly unseen during training) of a known given class and signal task completion when it determines it has reached the goal.
Implementation#
For this tutorial, we'll use the readily available objectnav_baselines project, which includes configurations for
a wide variety of object navigation experiments for both iTHOR and RoboTHOR. Since those configuration files are
defined for a single-node setup, we will mainly focus on the changes required in the machine_params and
training_pipeline methods.
Note that, in order to use the headless version of AI2-THOR, we currently need to install a specific THOR commit,
different from the default one in robothor_plugin. Note that this command is included in the configuration script
below, so we don't need to run this:
pip install --extra-index-url https://ai2thor-pypi.allenai.org ai2thor==0+91139c909576f3bf95a187c5b02c6fd455d06b48
The experiment config starts as follows:
import math
from typing import Optional, Sequence
import torch
import torch.optim as optim
from torch.optim.lr_scheduler import LambdaLR
from allenact.algorithms.onpolicy_sync.losses import PPO
from allenact.algorithms.onpolicy_sync.losses.ppo import PPOConfig
from projects.objectnav_baselines.experiments.robothor.objectnav_robothor_rgb_resnetgru_ddppo import (
ObjectNavRoboThorRGBPPOExperimentConfig as BaseConfig,
)
from allenact.utils.experiment_utils import (
Builder,
LinearDecay,
MultiLinearDecay,
TrainingPipeline,
PipelineStage,
)
class DistributedObjectNavRoboThorRGBPPOExperimentConfig(BaseConfig):
def tag(self) -> str:
return "DistributedObjectNavRoboThorRGBPPO"
We override ObjectNavRoboThorBaseConfig's THOR_COMMIT_ID to match the installed headless one:
THOR_COMMIT_ID = "91139c909576f3bf95a187c5b02c6fd455d06b48"
Also indicate that we're using headless THOR (for task_sampler_args methods):
THOR_IS_HEADLESS = True
Temporary hack Disable the commit_id argument passed to the THOR Controller's init method:
def env_args(self):
res = super().env_args()
res.pop("commit_id", None)
return res
And, of course, define the number of nodes. This will be used by machine_params and training_pipeline below.
We override the existing ExperimentConfig's init method to include control on the number of nodes:
def __init__(
self,
distributed_nodes: int = 1,
num_train_processes: Optional[int] = None,
train_gpu_ids: Optional[Sequence[int]] = None,
val_gpu_ids: Optional[Sequence[int]] = None,
test_gpu_ids: Optional[Sequence[int]] = None,
):
super().__init__(num_train_processes, train_gpu_ids, val_gpu_ids, test_gpu_ids)
self.distributed_nodes = distributed_nodes
Machine parameters#
Note: We assume that all nodes are identical (same number and model of GPUs and drivers).
The machine_params method will be invoked by runner.py with different arguments, e.g. to determine the
configuration for validation or training.
When working in distributed settings, AllenAct needs to know the total number of trainers across all nodes as well
as the local number of trainers. This is accomplished through the introduction of a machine_id keyword argument,
which will be used to define the training parameters as follows:
def machine_params(self, mode="train", **kwargs):
params = super().machine_params(mode, **kwargs)
if mode == "train":
params.devices = params.devices * self.distributed_nodes
params.nprocesses = params.nprocesses * self.distributed_nodes
params.sampler_devices = params.sampler_devices * self.distributed_nodes
if "machine_id" in kwargs:
machine_id = kwargs["machine_id"]
assert (
0 <= machine_id < self.distributed_nodes
), f"machine_id {machine_id} out of range [0, {self.distributed_nodes - 1}]"
local_worker_ids = list(
range(
len(self.train_gpu_ids) * machine_id,
len(self.train_gpu_ids) * (machine_id + 1),
)
)
params.set_local_worker_ids(local_worker_ids)
# Confirm we're setting up train params nicely:
print(
f"devices {params.devices}"
f"\nnprocesses {params.nprocesses}"
f"\nsampler_devices {params.sampler_devices}"
f"\nlocal_worker_ids {params.local_worker_ids}"
)
elif mode == "valid":
# Use all GPUs at their maximum capacity for training
# (you may run validation in a separate machine)
params.nprocesses = (0,)
return params
In summary, we need to specify which indices in devices, nprocesses and sampler_devices correspond to the
local machine_id node (whenever a machine_id is given as a keyword argument), otherwise we specify the global
configuration.
Training pipeline#
In preliminary ObjectNav experiments, we observe that small batches are useful during the initial training steps in terms of sample efficiency, whereas large batches are preferred during the rest of training.
In order to scale to the larger amount of collected data in multi-node settings, we will proceed with a two-stage pipeline:
- In the first stage, we'll enforce a number of updates per amount of collected data similar to the configuration with a single node by enforcing more batches per rollout (for about 30 million steps).
- In the second stage we'll switch to a configuration with larger learning rate and batch size to be used up to the grand total of 300 million experience steps.
We first define a helper method to generate a learning rate curve with decay for each stage:
@staticmethod
def lr_scheduler(small_batch_steps, transition_steps, ppo_steps, lr_scaling):
safe_small_batch_steps = int(small_batch_steps * 1.02)
large_batch_and_lr_steps = ppo_steps - safe_small_batch_steps - transition_steps
# Learning rate after small batch steps (assuming decay to 0)
break1 = 1.0 - safe_small_batch_steps / ppo_steps
# Initial learning rate for large batch (after transition from initial to large learning rate)
break2 = lr_scaling * (
1.0 - (safe_small_batch_steps + transition_steps) / ppo_steps
)
return MultiLinearDecay(
[
# Base learning rate phase for small batch (with linear decay towards 0)
LinearDecay(steps=safe_small_batch_steps, startp=1.0, endp=break1,),
# Allow the optimizer to adapt its statistics to the changes with a larger learning rate
LinearDecay(steps=transition_steps, startp=break1, endp=break2,),
# Scaled learning rate phase for large batch (with linear decay towards 0)
LinearDecay(steps=large_batch_and_lr_steps, startp=break2, endp=0,),
]
)
The training pipeline looks like:
def training_pipeline(self, **kwargs):
# These params are identical to the baseline configuration for 60 samplers (1 machine)
ppo_steps = int(300e6)
lr = 3e-4
num_mini_batch = 1
update_repeats = 4
num_steps = 128
save_interval = 5000000
log_interval = 10000 if torch.cuda.is_available() else 1
gamma = 0.99
use_gae = True
gae_lambda = 0.95
max_grad_norm = 0.5
# We add 30 million steps for small batch learning
small_batch_steps = int(30e6)
# And a short transition phase towards large learning rate
# (see comment in the `lr_scheduler` helper method
transition_steps = int(2 / 3 * self.distributed_nodes * 1e6)
# Find exact number of samplers per GPU
assert (
self.num_train_processes % len(self.train_gpu_ids) == 0
), "Expected uniform number of samplers per GPU"
samplers_per_gpu = self.num_train_processes // len(self.train_gpu_ids)
# Multiply num_mini_batch by the largest divisor of
# samplers_per_gpu to keep all batches of same size:
num_mini_batch_multiplier = [
i
for i in reversed(
range(1, min(samplers_per_gpu // 2, self.distributed_nodes) + 1)
)
if samplers_per_gpu % i == 0
][0]
# Multiply update_repeats so that the product of this factor and
# num_mini_batch_multiplier is >= self.distributed_nodes:
update_repeats_multiplier = int(
math.ceil(self.distributed_nodes / num_mini_batch_multiplier)
)
return TrainingPipeline(
save_interval=save_interval,
metric_accumulate_interval=log_interval,
optimizer_builder=Builder(optim.Adam, dict(lr=lr)),
num_mini_batch=num_mini_batch,
update_repeats=update_repeats,
max_grad_norm=max_grad_norm,
num_steps=num_steps,
named_losses={"ppo_loss": PPO(**PPOConfig, show_ratios=False)},
gamma=gamma,
use_gae=use_gae,
gae_lambda=gae_lambda,
advance_scene_rollout_period=self.ADVANCE_SCENE_ROLLOUT_PERIOD,
pipeline_stages=[
# We increase the number of batches for the first stage to reach an
# equivalent number of updates per collected rollout data as in the
# 1 node/60 samplers setting
PipelineStage(
loss_names=["ppo_loss"],
max_stage_steps=small_batch_steps,
num_mini_batch=num_mini_batch * num_mini_batch_multiplier,
update_repeats=update_repeats * update_repeats_multiplier,
),
# The we proceed with the base configuration (leading to larger
# batches due to the increased number of samplers)
PipelineStage(
loss_names=["ppo_loss"],
max_stage_steps=ppo_steps - small_batch_steps,
),
],
# We use the MultiLinearDecay curve defined by the helper function,
# setting the learning rate scaling as the square root of the number
# of nodes. Linear scaling might also works, but we leave that
# check to the reader.
lr_scheduler_builder=Builder(
LambdaLR,
{
"lr_lambda": self.lr_scheduler(
small_batch_steps=small_batch_steps,
transition_steps=transition_steps,
ppo_steps=ppo_steps,
lr_scaling=math.sqrt(self.distributed_nodes),
)
},
),
)
Multi-node configuration#
Note: In the following, we'll assume you don't have an available setup for distributed execution, such as slurm. If you do have access to a better alternative to setup and run distributed processes, we encourage you to use that. The experimental distributed tools included here are intended for a rather basic usage pattern that might not suit your needs.
If we haven't set up AllenAct with the headless version of Ai2-THOR in our nodes, we can define a configuration script similar to:
#!/bin/bash
# Prepare a virtualenv for allenact
sudo apt-get install -y python3-venv
python3 -mvenv ~/allenact_venv
source ~/allenact_venv/bin/activate
pip install -U pip wheel
# Install AllenAct
cd ~
git clone https://github.com/allenai/allenact.git
cd allenact
# Install AllenaAct + RoboTHOR plugin dependencies
pip install -r requirements.txt
pip install -r allenact_plugins/robothor_plugin/extra_requirements.txt
# Download + setup datasets
bash datasets/download_navigation_datasets.sh robothor-objectnav
# Install headless AI2-THOR and required libvulkan1
sudo apt-get install -y libvulkan1
pip install --extra-index-url https://ai2thor-pypi.allenai.org ai2thor==0+91139c909576f3bf95a187c5b02c6fd455d06b48
# Download AI2-THOR binaries
python -c "from ai2thor.controller import Controller; c=Controller(); c.stop()"
echo DONE
and save it as headless_robothor_config.sh. Note that some of the configuration steps in the script assume you have
superuser privileges.
Then, we can just copy this file to the first node in our cluster and run it with:
source <PATH/TO/headless_robothor_config.sh>
If everything went well, we should be able to
cd ~/allenact && source ~/allenact_venv/bin/activate
Note that we might need to install libvulkan1 in each node (even if the AllenAct setup is shared across nodes) if it
is not already available.
Local filesystems#
If our cluster does not use a shared filesystem, we'll need to propagate the setup to the rest of nodes. Assuming
we can just ssh with the current user to all nodes, we can propagate our config with
scripts/dconfig.py --runs_on <COMMA_SEPARATED_LIST_OF_IP_ADDRESSES> \
--config_script <PATH/TO/headless_robothor_config.sh>
and we can check the state of the installation with the scripts/dcommand.py tool:
scripts/dcommand.py --runs_on <COMMA_SEPARATED_LIST_OF_IP_ADDRESSES> \
--command 'tail -n 5 ~/log_allenact_distributed_config'
If everything went fine, all requirements are ready to start running our experiment.
Run your experiment#
Note: In this section, we again assume you don't have an available setup for distributed execution, such as slurm. If you do have access to a better alternative to setup/run distributed processes, we encourage you to use that. The experimental distributed tools included here are intended for a rather basic usage pattern that might not suit your needs.
Our experimental extension to AllenAct's main.py script allows using practically identical commands to the ones
used in a single-node setup to start our experiments. From the root allenact directory, we can simply invoke
scripts/dmain.py projects/tutorials/distributed_objectnav_tutorial.py \
--config_kwargs '{"distributed_nodes":3}' \
--runs_on <COMMA_SEPARATED_LIST_OF_IP_ADDRESSES> \
--env_activate_path ~/allenact_venv/bin/activate \
--allenact_path ~/allenact \
--distributed_ip_and_port <FIRST_IP_ADDRESS_IN_RUNS_ON_LIST>:<FREE_PORT_NUMBER_FOR_THIS_IP_ADDRESS>
This script will do several things for you, including synchronization of the changes in the allenact directory
to all machines, enabling virtual environments in each node, sharing the same random seed for all main.py instances,
assigning --machine_id parameters required for multi-node training, and redirecting the process output to a log file
under the output results folder.
Note that by changing the value associated with the distributed_nodes key in the config_kwargs map and the runs_on
list of IPs, we can easily scale our training to e.g. 1, 3, or 8 nodes as shown in the chart above. Note that for this
call to work unmodified, you should have sufficient GPUs/GPU memory to host 60 samplers per node.
Track and stop your experiment#
You might have noticed that, when your experiment started with the above command, a file was created under
~/.allenact. This file includes IP addresses and screen session IDs for all nodes. It can be used
by the already introduced scripts/dcommand.py script, if we omit the --runs_on argument, to call a command on each
node via ssh; but most importantly it is used by the scripts/dkill.py script to terminate all screen sessions hosting
our training processes.
Experiment tracking#
A simple way to check all machines are training, assuming you have nvidia-smi installed in all nodes, is to just call
scripts/dcommand.py
from the root allenact directory. If everything is working well, the GPU usage stats from nvidia-smi should reflect
ongoing activity. You can also add different commands to be executed by each node. It is of course also possible to run
tensorboard on any of the nodes, if that's your preference.
Experiment termination#
Just call
scripts/dkill.py
After killing all involved screen sessions, you will be asked about whether you also want to delete the "killfile"
stored under the ~/.allenact directory (which might be your preferred option once all processes are terminated).
We hope this tutorial will help you start quickly testing new ideas! Even if we've only explored moderates settings of up to 480 experience samplers, you might want to consider some additional changes (like the choice for the optimizer) if you plan to run at larger scale.